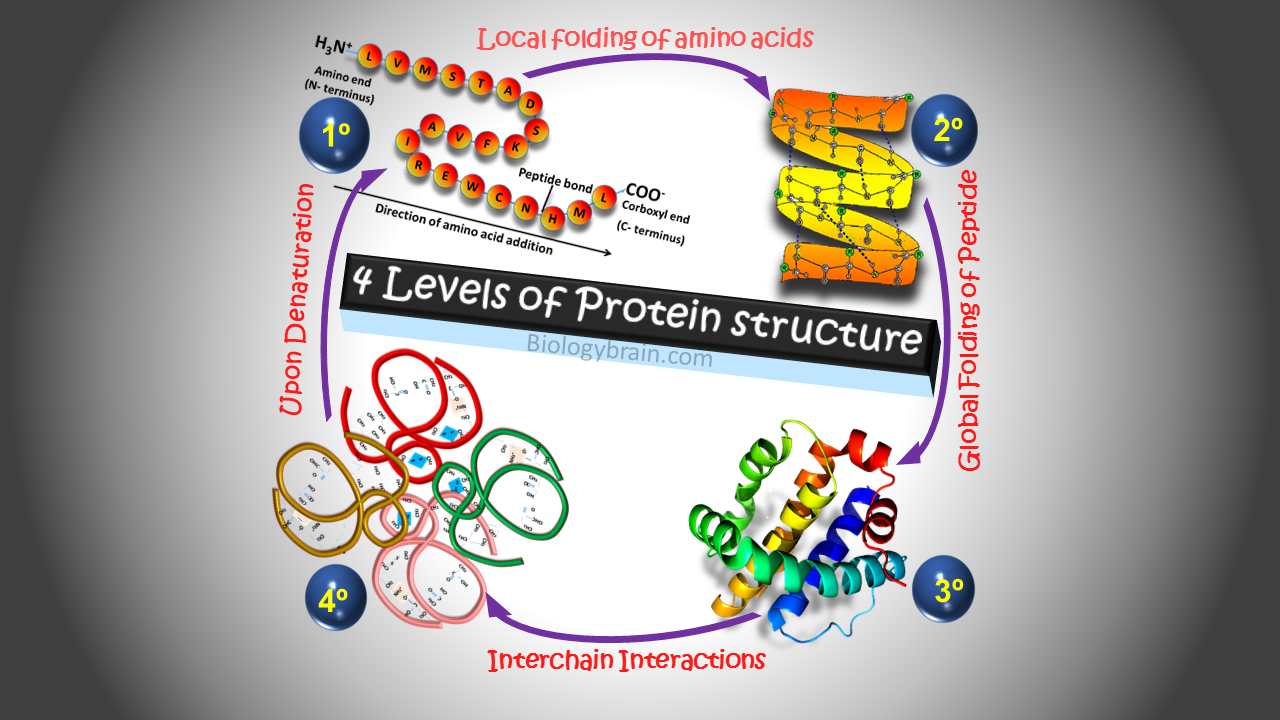
The term structure when used in the relation to proteins, takes on a much more complex meaning than it does for small molecules. Proteins are macromolecules and have four different levels of structures – Primary, Secondary, Tertiary, and Quaternary.
1) Primary structure: It represents the linear amino acid structure
It is a basic level of protein structure, which is a simple linear sequence of amino acids that form a polypeptide chain without any intrachain interactions.
2) Secondary structure: Local folding of amino acids found in the primary structure.
Examples: α-helix contained protein structure including α-keratin, myoglobin, and hemoglobin.
While the commercially valuable insoluble protein of silk such as fibroin and sericin, etc. are antiparallel β-pleated sheet-containing proteins.
3) Tertiary structure: Global folding of peptide strand.
Examples: The albumin, insulin, and monomeric form of actin are the best example of globular proteins.
4) Quaternary structure: Interactions between two subunits lead to the binding of two subunits and form the functional proteins.
Examples: Cro protein (DNA-binding protein) of bacteriophage λ, hemoglobin, collagen, Insulin, myoglobin, etc.
Changes in the protein-coding sequence of a gene can have the following effects
1) Silent mutation: This occurs when one codon is replaced by another codon that can specify (coding) the same amino acid. This replacement will not change the amino acid sequence of the protein, so the function of the protein will be remaining the same.
2) Missense substitution (point mutation): In this substitution, a sense codon specifying a particular amino acid is substituted by a sense codon specifying other amino acids. This substitution can change the amino acid sequence of the protein.
3) Nonsense substitution: In this substitution, a sense codon specifying an amino acid is substituted by termination or stop codon that results in premature termination of translation. This mutation will lead to the formation of truncated protein. Truncated proteins may behave like uncontrollable elements.
Research scope on protein structure:
The present studies are focusing mainly on “drug-target interactions”, especially proteins as therapeutic targets. Production of protein formulations can be a major challenge. Gathering and providing information related to proteins and their four structural levels may be the concerned strategies to analyze drug-protein interactions. However, without good knowledge of the nature of the protein structures and the conformational characteristics of the specific protein, the development of potential drugs against the target proteins will be a very difficult task.
This brief technical information offers researchers get a quick overview of protein structure and its mechanism of action. In this article, we are providing brief information on the different structures of proteins. It will also cover how protein structures can be affected during the formulation and some of the analytical methods which can be used to determine the protein structure and analyze the protein stability.
Determination of amino acid sequence in a protein can be used to identify the primary structure of the protein. It is the only structure that shows higher stabilization by covalent interactions at the structural level. However, the amino acid sequence at the primary structural level does not contain non-covalent interactions. The covalent interactions of the peptides present between two amino acids. The primary structure will be formed by covalent interactions between α- carboxylic and α- amino groups of the amino acids. The side chains of the amino acids do not participate in the primary structure formation.